Introduction¶
This package provides Python implementation of Augmented Principal Component Analysis (AugmentedPCA) models - a family of linear factor models that find a set of factors aligned with an augmenting objective in addition to the canonical PCA objective of finding factors that represent the data variance. AugmentedPCA models can be split into two general families of models: adversarial AugmentedPCA and supervised AugmentedPCA.
Models Overview¶
AugmentedPCA has two main model variants: adversarial AugmentedPCA (AAPCA) and supervised AugmentedPCA (SAPCA). A brief introduction to the two variants is given in the following sections.
Supervised AugmentedPCA¶
In supervised AugmentedPCA (SAPCA), the augmenting objective is to make the factors aligned with the data labels, or some outcome, in addition to having the factors explain the variance of the original observed or primary data. SAPCA is useful when predictivity of latent components with respects to a set of data labels or outcomes is desired. SAPCA is equivalent to a supervised autoencoder (SAE) with a single hidden layer. Therefore, SAPCA can be applied to situations where the properties of latent representations enforced via deep SAEs are desired, yet where limited data or training inconsistencies are a concern. Below is a diagram depicting the relationship between primary data, supervision data, and the resulting SAPCA factors.
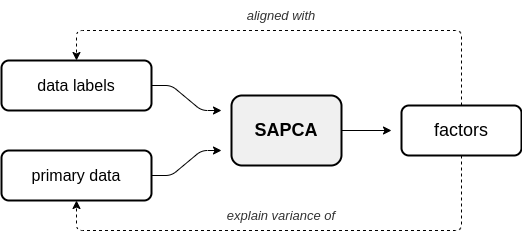
Adversarial AugmentedPCA¶
In adversarial AugmentedPCA (AAPCA), the augmenting objective is to make the factors orthogonal to a set of concomitant data, in addition to having the factors explain the variance of the original observed or primary data. AAPCA can be used in situations where one wishes to enforce invariance of latent components to a set of concomitant data, and is equivalent to an adversarial autoencoder with a single hidden layer. Below is a diagram depicting the relationship between primary data, concomitant data, and the resulting AAPCA factors.
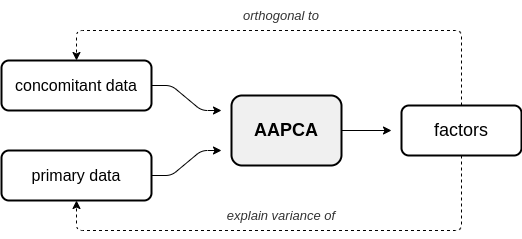
Quick Introduction¶
Below is quick guide to using AugmentedPCA. For a more in-depth examples, see the Examples section of the documentation.
Importing AugmentedPCA Models¶
AugmentedPCA models can be imported from the models.py module. Below we show an example of importing the
AAPCA model.
# Import all AugmentedPCA models
from apca.models import AAPCA
Alternatively, all offered AugmentedPCA models can be imported at once.
# Import all AugmentedPCA models
from apca.models import *
Instantiating AugmentedPCA Models¶
AugmentedPCA models are instantiated by assigning either an SAPCA or AAPCA object to a variable. During instantiation,
one has the option to define parameters n_components, mu, which represent the number of components
and the augmenting objective strength, respectively. Additionally, approximate inference strategy can be defined
through the inference parameter.
# Define model parameters
n_components = 2 # factors will have dimensionality of 2
mu = 1.0 # augmenting objective strength equal to 1
inference = 'encoded' # encoded approximate inference strategy
# Instantiate adversarial AugmentedPCA model
aapca = AAPCA(n_components=n_components, mu=mu, inference=inference)
Fitting APCA¶
AugmentedPCA models closely follow the style and implemention of scikit-learn’s PCA implementation, with many of the same methods and
functionality. Similar to scikit-learn models, AugmentedPCA models are fit using the fit() method.
fit() takes two parameters: X which represents the matrix of primary data and Y which
represents the matrix of augmenting data.
Note
Before fitting AugmentedPCA models, it may be helpful to scale both the primary and augmenting data. Having the
primary and augmenting data on the same scale will result in more consistent range of effective augmenting
objective strengths (controlled by the mu paramter) across different datasets.
# Import numpy
import numpy as np
# Generate synthetic data
# Note: primary and augmenting data must have same number of samples/same first dimension size
n_samp = 100
X = np.random.randn(n_samp, 20) # primary data, 100 samples with dimensionality of 20
Y = np.random.randn(n_samp, 3) # concomitant data, 100 samples with dimensionality of 3
# Fit adversarial AugmentedPCA model
aapca.fit(X=X, Y=Y)
Alternatively, AugmentedPCA models can be fit using the fit_transform() method, which takes the same
parameters as the fit() method but also returns a matrix of components or factors.
# Fit adversarial AugmentedPCA model and generate components
S = aapca.fit_transform(X=X, Y=Y)